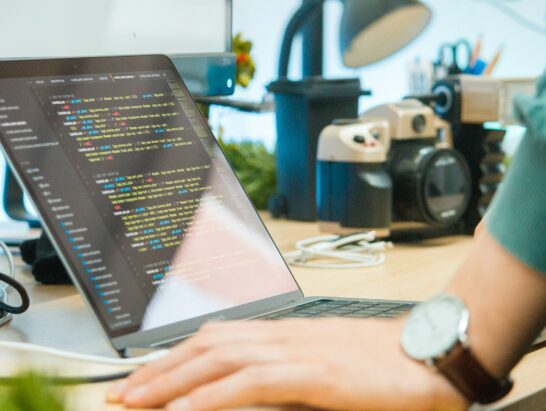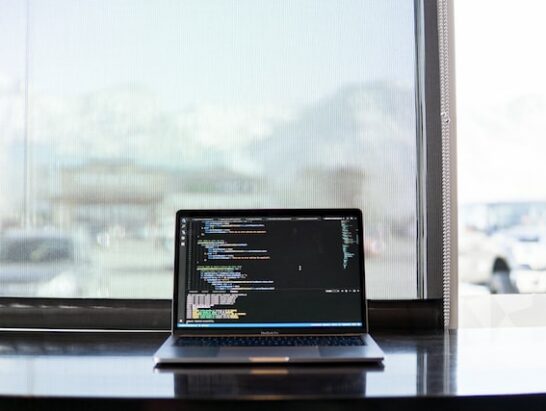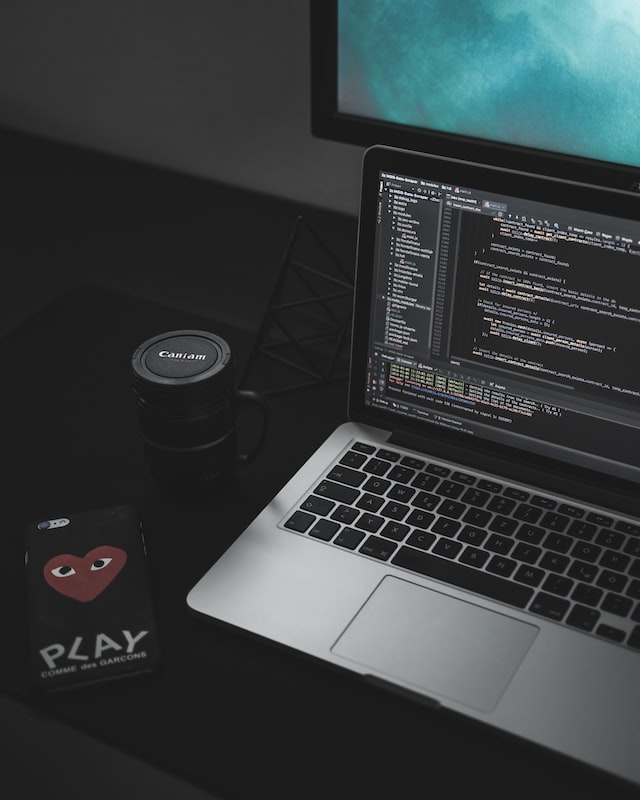
Online Conference
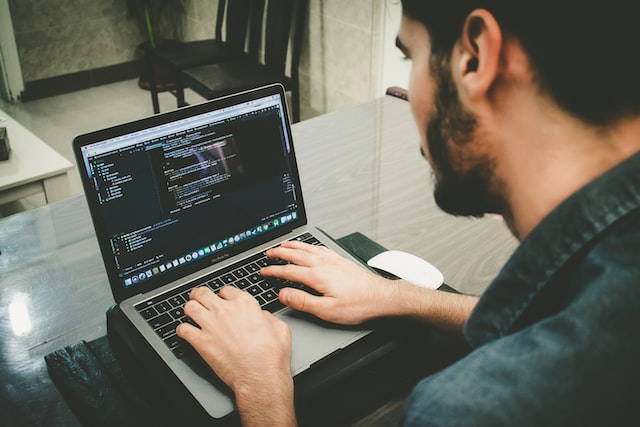

working experience

Online conference for Middle professionals
- World Leaders The world leaders of IT and Digital area, representatives of major companies, leading experts from the UK, USA and Europe.
- Trends and Tools Current trends and tools that will be the first confident step to successful employment in high-paying fields.
Registration and pre-moderation are required for free participation
Awards and Certifications

Play at a Finnish casino without registration or casino ei rekisteröintiä, as it’s called locally.

Our partners from CasinoHEX Philippines prepared the review of 22be, new online casino for the Philippines market. Gamblers can explore 22bet casino Philippines bonus offers, latest slots collection and betting app review.


IT college students have a lot on their plates, leaving little time to attend FP REAPIT programming conference. Going online for homework help with programming assignments when you’re stuck on a coding assignment may save you hours of sitting at the computer and free your time for essential events.

Are you a future programming pro? Or are you a top-notch professional with solid conference experience who wants to share your insights? Anyway, you should go on Twitter, both an entertaining and educational platform. And if you want to deliver your messages to the audience more effectively, you should buy likes on Twitter. Purchase real Twitter likes for cheap from the best UK provider and boost your professional reputation online!



SocoLive.Soccer brings the football stadium atmosphere right to your living room.

Find the best online casino in Australia to play for real money: Aucasinoonline.com
Latest From Our Conference
Development in the cloud
Why learn new programming languages
Review of the new version of Swift 5.5
Mobile app on Flutter: pros and cons for business
Online conference about programming
Testimonials
Mahfuz Riad
Mahfuz Riad
Mahfuz Riad